If the cells you would like to access are currently listed as unavailable
or you are ordering from outside of Europe please get in touch via
Contact@EBiSC.org.
UKBi005-A
iLB-C-31f-r1, LB-31-1
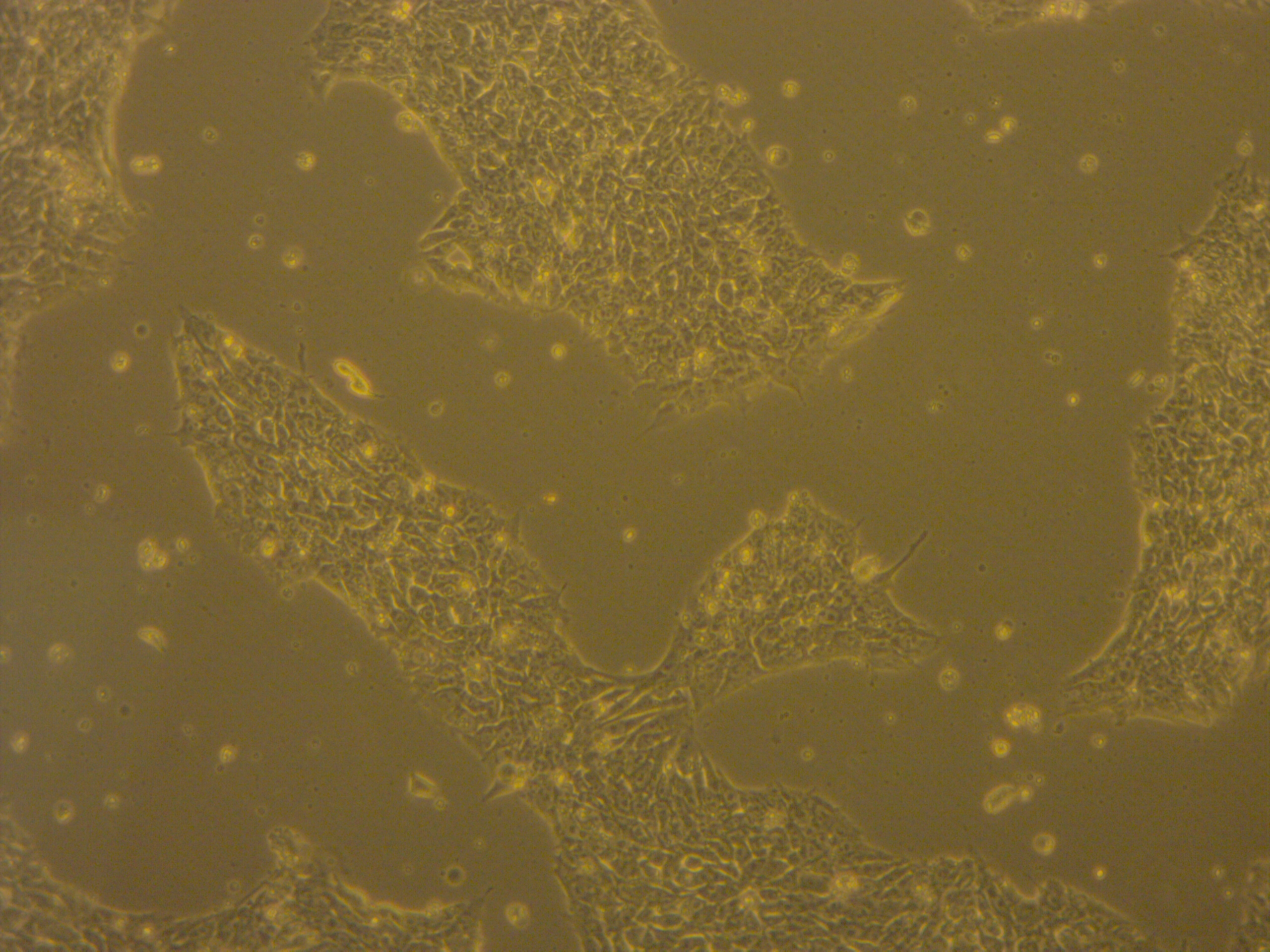
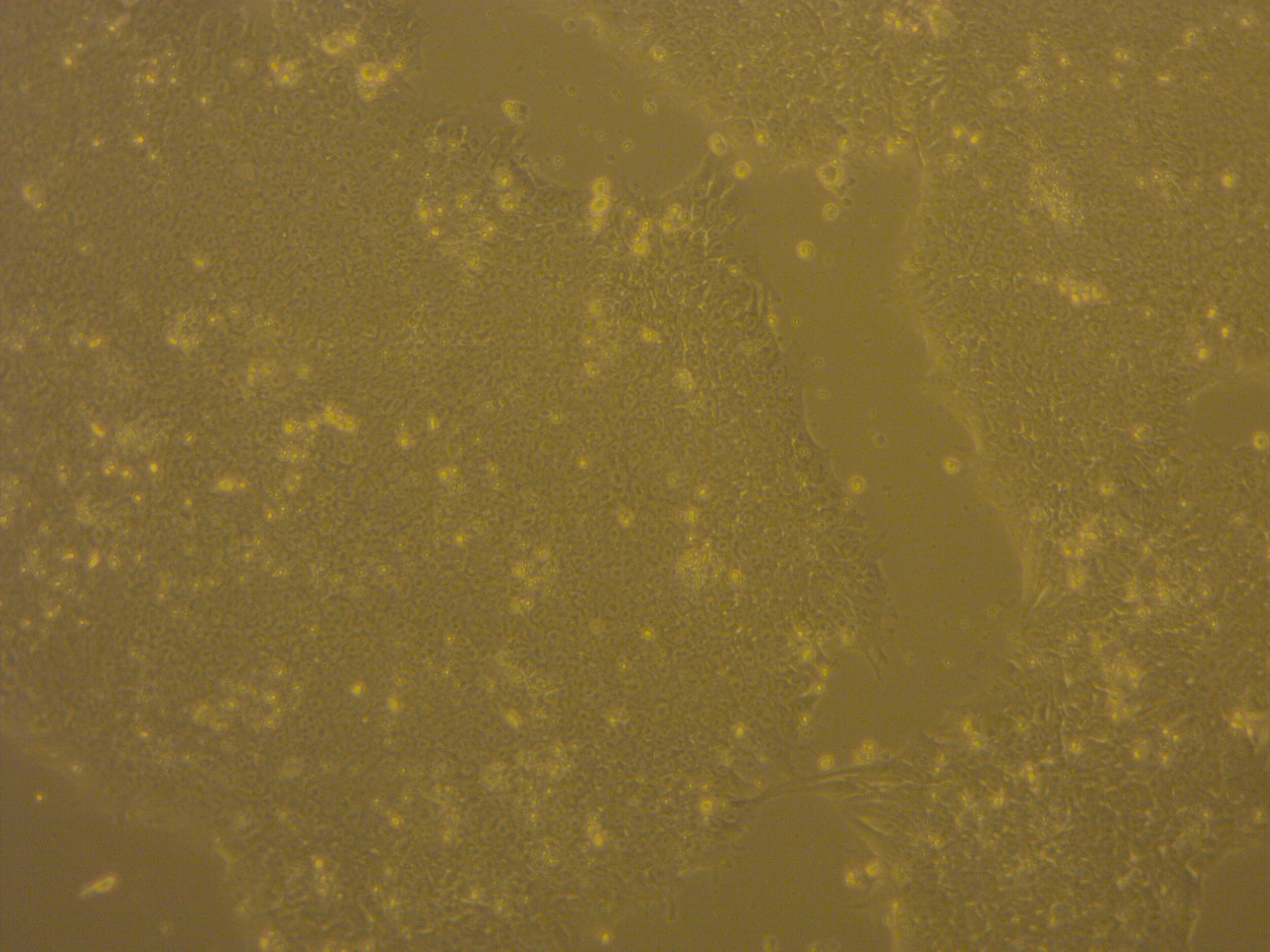
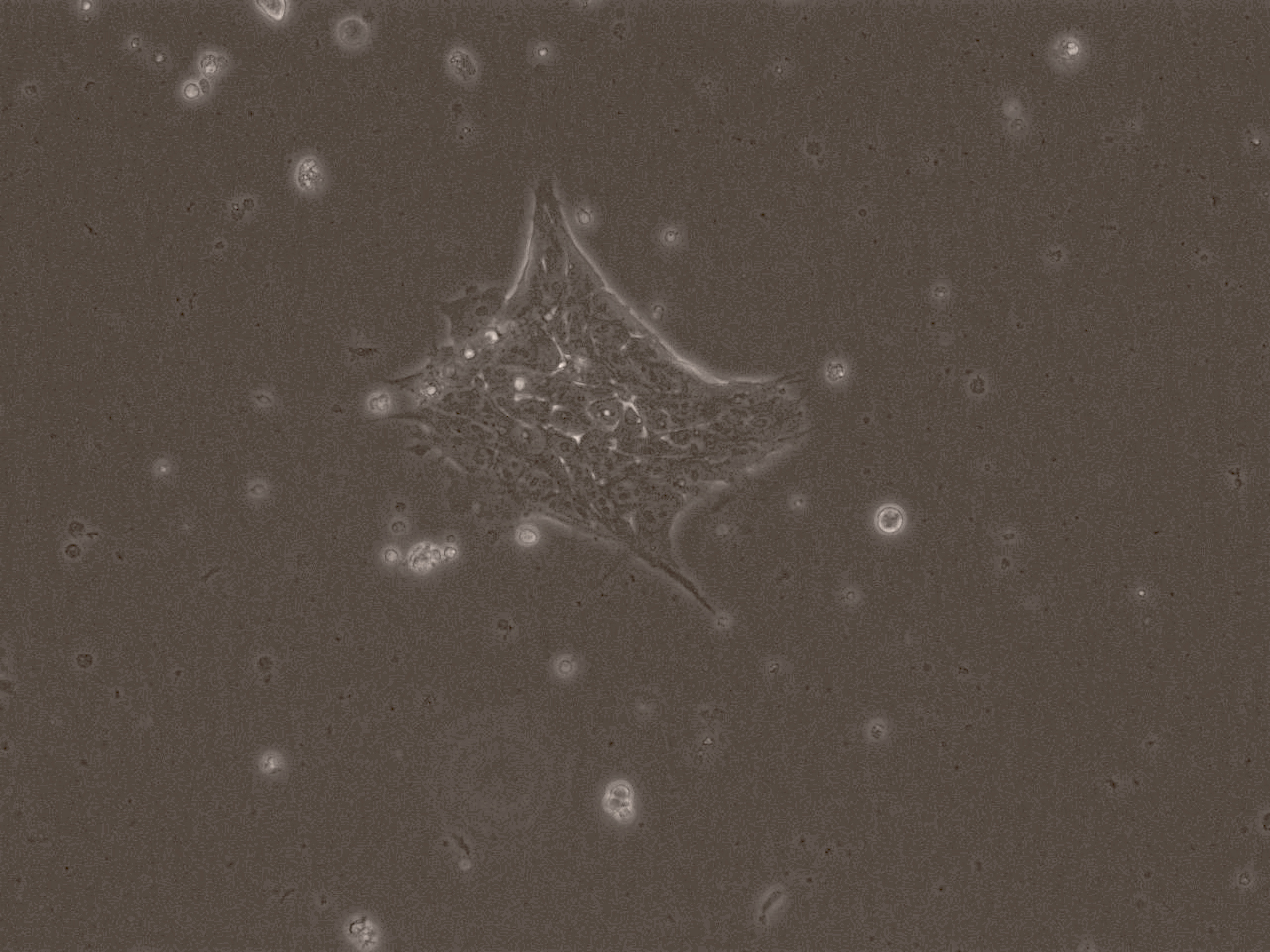
iPSC line
A CLIP contains information about a cell line including any
specific third party obligations relating to, for example,
licensing obligations or the donor consent which affect the
use of the cell line.
The EBiSC Access and Use Agreement must be completed along with an individual
Cell Line Information Pack for each line. Complete the EAUA and send to Contact@EBiSC.org
for countersignature. The EAUA must be fully signed before proceeding with your order.
A batch specific Certificate of Analysis will be available to
download once you receive your EBiSC iPSC line.
General#
Cell Line |
|
| hPSCreg name | UKBi005-A |
| Alternative name(s) |
iLB-C-31f-r1, LB-31-1
|
| Cell line type | Human induced pluripotent stem cell (hiPSC) |
| Similar lines | No similar lines found. |
Provider |
|
| Depositor | Universitätsklinikum Bonn (UKB) |
| Owner | Universitätsklinikum Bonn |
| Distributors |
EBiSC
Institut für Rekonstruktive Neurobiologie
Scottish Centre for Regenerative Medicine
|
| Derivation country | Germany |
External Databases |
|
| hPSCreg | UKBi005-A |
| BioSamples | SAMEA4584351 |
| Cellosaurus | CVCL_1E85 |
| Wikidata | Q54990251 |
General Information |
|
| Publications | View all related publications on hPSCreg (4) |
| This EBiSC line can be used for: |
Yes
Research use: allowed
Clinical use: no
Commercial use: no
|
Donor Information#
General Donor Information |
|
| Sex | female |
| Age of donor (at collection) | 20-24 |
Phenotype and Disease related information (Donor) |
|
| Diseases | No disease was diagnosed.
|
External Databases (Donor) |
|
| BioSamples | SAMEA4584241 |
hIPSC Derivation#
General |
|
| Source cell type | |
| Source cell origin |
Any portion of the organ that covers that body and consists of a layer of epidermis and a layer of dermis.
Synonyms
|
| Age of donor (at collection) | 20-24 |
| Collected in | 2009 |
| Passage number reprogrammed | P4 |
Reprogramming method |
|
| Vector type | Integrating |
| Vector | Virus (Retrovirus) |
| Genes | |
| Is the used vector excisable? |
No |
| Absence of reprogramming vector(s)? |
Yes |
| Reprogramming vectors silenced? |
Yes |
| Methods used |
PCR
|
| Files and images showing reprogramming vector expressed or silenced | |
Vector free reprogramming |
|
| Type of used vector free reprogramming factor(s) |
None
|
Other |
|
| Selection criteria for clones | Morphology |
| Derived under xeno-free conditions |
No |
| Derived under GMP? |
No |
| Available as clinical grade? |
No |
Culture Conditions#
Latest released batch |
|
| Culture medium | mTeSR |
| Passage method | EDTA |
| Surface coating | Matrigel / Geltrex |
| O2 concentration | 21 |
| CO2 concentration | 5 |
| Temperature | 37 |
The following are the depositor culture conditions, they do not refer to any specific batch.
| Surface coating | Matrigel/Geltrex |
| Feeder cells |
No |
| Passage method |
Enzyme-free cell dissociation
EDTA
|
| O2 Concentration | 21 % |
| CO2 Concentration | 5 % |
| Medium |
mTeSR™ 1
|
| Has Rock inhibitor (Y27632) been used at passage previously with this cell line? | No |
| Has Rock inhibitor (Y27632) been used at cryo previously with this cell line? | No |
| Has Rock inhibitor (Y27632) been used at thaw previously with this cell line? | No |
Characterisation#
Analysis of Undifferentiated Cells
| Marker | Expressed | Immunostaining | RT-PCR | Flow Cytometry | Enzymatic Assay | Expression Profiles |
| POU5F1 (OCT-4) |
Yes |
|||||
| SSEA-3 |
Yes |
|||||
| TRA 1-60 |
Yes |
|||||
| TRA 1-81 |
Yes |
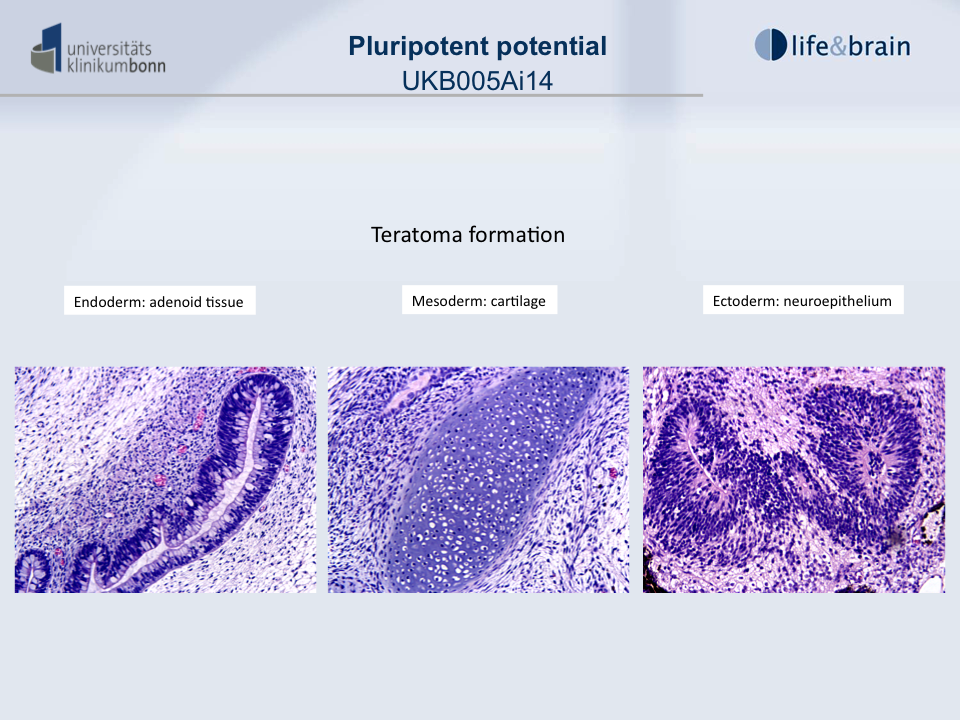
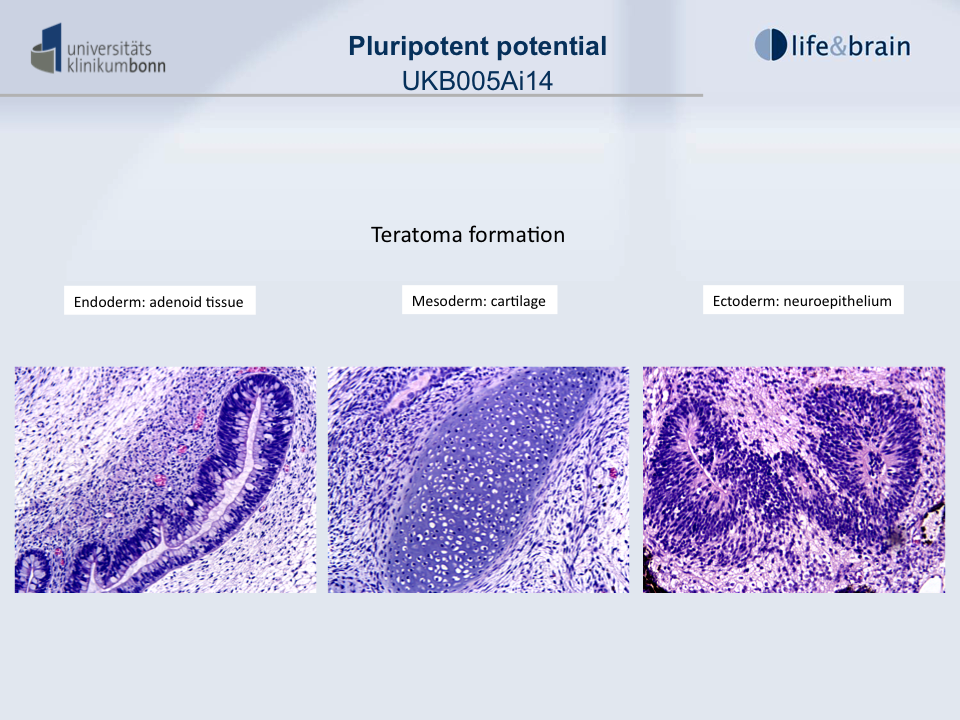
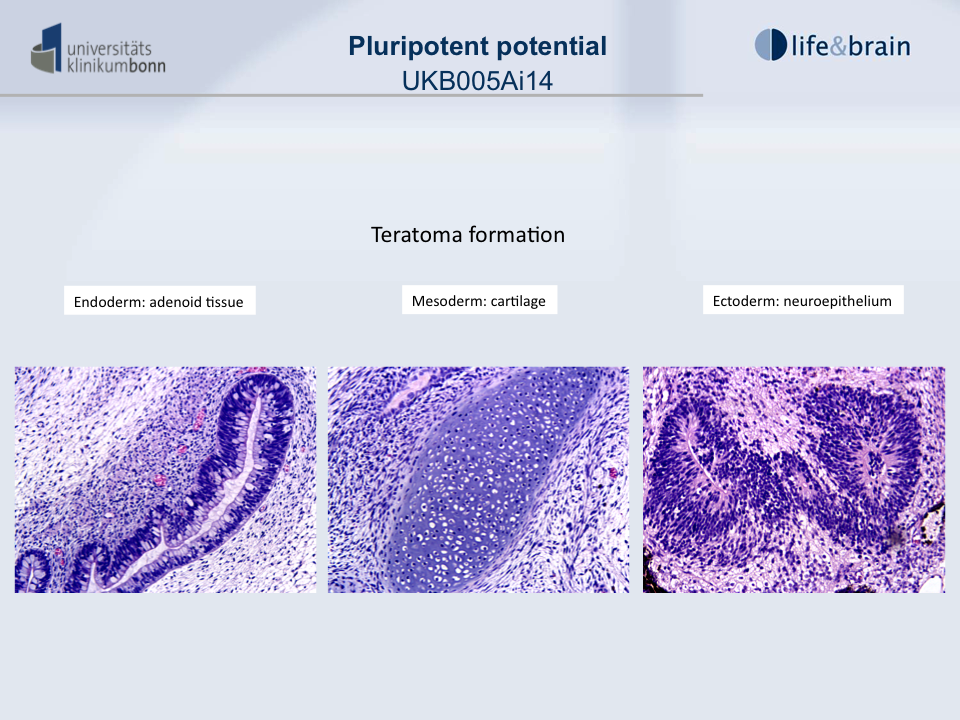
Differentiation Potency
In vivo teratoma
In vitro directed differentiation
Microbiology / Virus Screening |
|
| HIV 1 | Negative |
| HIV 2 | Negative |
| Hepatitis B | Negative |
| Hepatitis C | Negative |
| Mycoplasma | Negative |
Genotyping#
Karyotyping (Cell Line) |
|
| Has the cell line karyotype been analysed? |
Yes
|
Other Genotyping (Cell Line) |
|
| Is there genome-wide genotyping or functional data available? |
Yes
|